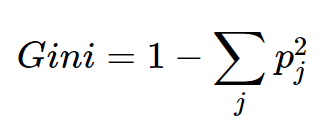- On each node, a decision tree asks a question and answers it with yes/no to classify observations under that node.
- The question could be based on a specific True/False (is obs.feature = something) question or could be based on numeric data (is obs.feature > 10)
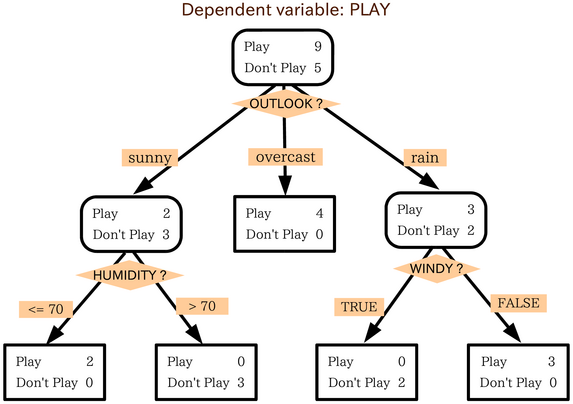
- The classification can be categorical or numerical
- The first node is called the Root node, the nodes in the middle are called internal nodes, and the results are called leaf nodes
In [44]:
# source https://scikit-learn.org/stable/modules/tree.html#classification
from sklearn import tree
from sklearn.datasets import load_iris
from sklearn import tree
import pandas as pd
iris = load_iris()
X, y = load_iris(return_X_y=True)
X_df = pd.DataFrame(X, columns=iris.feature_names)
X_df.head()
Out[44]:
| sepal length (cm) | sepal width (cm) | petal length (cm) | petal width (cm) | |
|---|---|---|---|---|
| 0 | 5.1 | 3.5 | 1.4 | 0.2 |
| 1 | 4.9 | 3.0 | 1.4 | 0.2 |
| 2 | 4.7 | 3.2 | 1.3 | 0.2 |
| 3 | 4.6 | 3.1 | 1.5 | 0.2 |
| 4 | 5.0 | 3.6 | 1.4 | 0.2 |
In [45]:
# Imagine we need to pick a root node ourselves: we'll need to see how each feature performs on the split
## Fitting our tree on the train data
import matplotlib.pyplot as plt
for feature in iris.feature_names:
plt.figure(figsize=(10,6))
clf = tree.DecisionTreeClassifier(max_depth=1)
clf = clf.fit(X_df.loc[:,feature].values.reshape(-1,1), y)
print("Decision tree based on {0}".format(feature))
tree.plot_tree(clf, filled=False, impurity=False)
plt.show()
Decision tree based on sepal length (cm)
Decision tree based on sepal width (cm)
Decision tree based on petal length (cm)
Decision tree based on petal width (cm)
- When the leaves don't contain 100% of our data, we consider them "impure"
- Now, to get our Root node, we'll need to measure this "impurity" concept and choose the "best" feature as a root
- The most popular way to measure impurity is Gini (more on it here interpretation and intuitive explanation of Gini impurity )
- So now, based on this impurity measure we can evaluate what the best split is (all we'll to do is calculate the weighted average for the leaf nodes of each split)
In [46]:
# This process is already implemented in the sklearn library.
## So let's look at what it yields!
clf = tree.DecisionTreeClassifier()
plt.figure(figsize=(20,10))
tree.plot_tree(clf.fit(X, y), impurity=True, feature_names=iris.feature_names)
plt.show()
In [47]:
## Plotting the tree (alternative - useful when tree is big!)
import graphviz
dot_data = tree.export_graphviz(clf, out_file=None)
graph = graphviz.Source(dot_data)
graph.render("iris")
dot_data = tree.export_graphviz(clf, out_file=None,
feature_names=iris.feature_names,
class_names=iris.target_names,
filled=True, rounded=True,
special_characters=True)
graph = graphviz.Source(dot_data)
graph
# see also: plot Decision surface of a decision tree using paired features
# https://scikit-learn.org/stable/auto_examples/tree/plot_iris_dtc.html
Out[47]: